Degradation of 2-chloroethanol by wild type and mutants of Pseudomonas putida US2
Time:2021/8/2 13:55:35
A strain of Pseudomonas putida was isolated that was able to degrade 2-chloroethanol. The degradation proceeded via 2-chloroacetaldehyde and chloroacetate to glycolate. In crude extracts the enzymes for this degradation pathway could be detected. All enzymes proved to be inducible. The dehalogenase that catalyzed the dehalogenation of chloroacetate to glycolate was further characterized. It consisted of a single polypeptide chain with a molecular mass of 28 kDa. After induction the dehalogenase was expressed at a high level. In a mutant resistant to high concentrations of 2-chloroethanol the dehalogenase was no longer expressed. The mechanism of resistance seemed to be due to the inability to convert chloroacetate and export of this compound out of the cell.
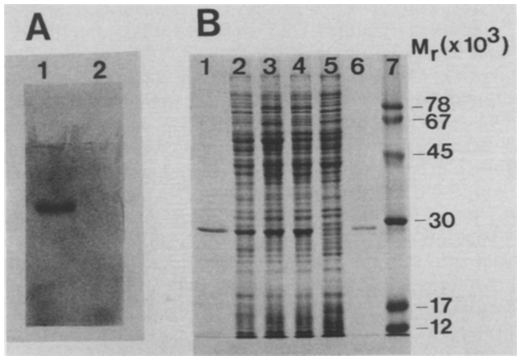
A Native polyacrylamide gele]eetrophoresis to visualize dehalogenase activity. Crude extracts were subjected to electrophoresis and dehalogenase was vis~xalized with an activity stain. Extracts were prepared from strain P. putida US2 grown on 2-chloroethanol (1) and Nutrient Broth (2). B SDS-polyacrylamide gelelectrophoresis of" crude extracts and purified dehaiogenase. Slots are: (1), (6), purified dchalogcnase, (2), (3), (4), crude extract of strain US2 grown on citrate, 2-chloroethanol and chloroacctate, respectively; (5), crude extract from mutant US2M1 grown on citrate; (7), marker proteins.
![]()
Regulation of PES-dcpendent chloroethanol dehydrogenase, NAD-dependent chloroaeetaldchyde dehydrogenase and chloroaeetate dehalogenase formation in Pseudomonas Imlidu.


 < /a>
< /a>